Mari Tinholt1,2, Nina Iversen1 and Per Morten Sandset2,3,*
1Department of Medical Genetics, Oslo University Hospital, Oslo, Norway.
2Department of Haematology, Oslo University Hospital, Oslo, Norway.
3Institute of Clinical Medicine, University of Oslo, Oslo, Norway.
The bidirectional link between cancer and thrombosis observed in epidemiological studies is well recognized. More recently, blood coagulation proteins have been proposed to play a role in cancer progression. Experimental data suggest that coagulation factors like tissue factor (TF), factor (F) VIIa, and FXa have tumor promoting effects [1-3], while the coagulation inhibitor TF pathway inhibitor (TFPI) exerts anti-tumor effects [4-6]. How the coagulation system relates to cancer etiology and risk at the genetic level is, however, largely unexplored.
The FV Leiden (rs6025) and the prothrombin gene G20210A (rs1799963) thrombophilic variants are the most studied polymorphisms in coagulation genes, but there is lack of consensus across studies. Only one cancer study has hitherto been large enough to study homozygous FV Leiden carriers [7], and this study found that homozygous carriers of FV Leiden had a 6-fold increased risk of colorectal cancer. No homozygous carriers of the prothrombin gene G20210A were observed. Heterozygosity for either the FV Leiden or the prothrombin gene G20210A variants was associated with a 30% reduced risk of colorectal cancer. Moreover, while another study reported a higher frequency of heterozygous carriers of the prothrombin gene G20210A variant in gastrointestinal cancer [8], most studies do not support a major role for FV Leiden or prothrombin G20210A heterozygosity as risk factors for solid cancer susceptibility [8-14].
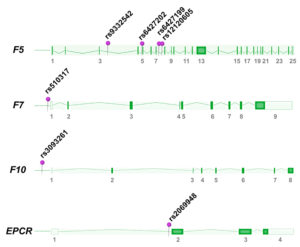

One Turkish study investigated selected polymorphisms of the F7 gene and demonstrated that the promoter SNP -402GA (rs510317) was associated with increased susceptibility of breast cancer [15]. In our Norwegian breast cancer study, we investigated 37 tagSNPs located in genes of the tissue factor pathway. We found that six SNPs in either the F5, F10, or EPCR genes were associated with breast cancer (F5 rs12120605, F5 rs6427202, F5 rs9332542, F5 rs6427199, F10 rs3093261, EPCR rs2069948) [16]), suggesting a possible role for these variants in breast cancer etiology. All F5 SNPs were independent of FV Leiden, and tended to be preferentially associated with hormone receptor negative and triple negative patients, thus reflecting the heterogeneity of breast cancer disease. The SNPs in our study could not explain systemic hypercoagulability (higher D-dimer levels, higher peak thrombin and increased activated protein C resistance), but could still contribute to the coagulation regulation within the tumor microenvironment. The F5 SNPs were later confirmed to regulate F5 expression in breast tumors (unpublished).
The differences in cancer susceptibility in the normal population are likely attributable to common genetic variation, of which SNPs are the most common. Genome-wide and candidate gene association studies revealing putative risk loci have increased our biological insights into carcinogenesis. However, the individual risk contribution of SNPs is typically small and SNPs may be indirectly linked to the true causal variant(s) through linkage disequilibrium rather than being the causal variant itself. The interpretation of the results from genetic association studies have therefore been challenging.
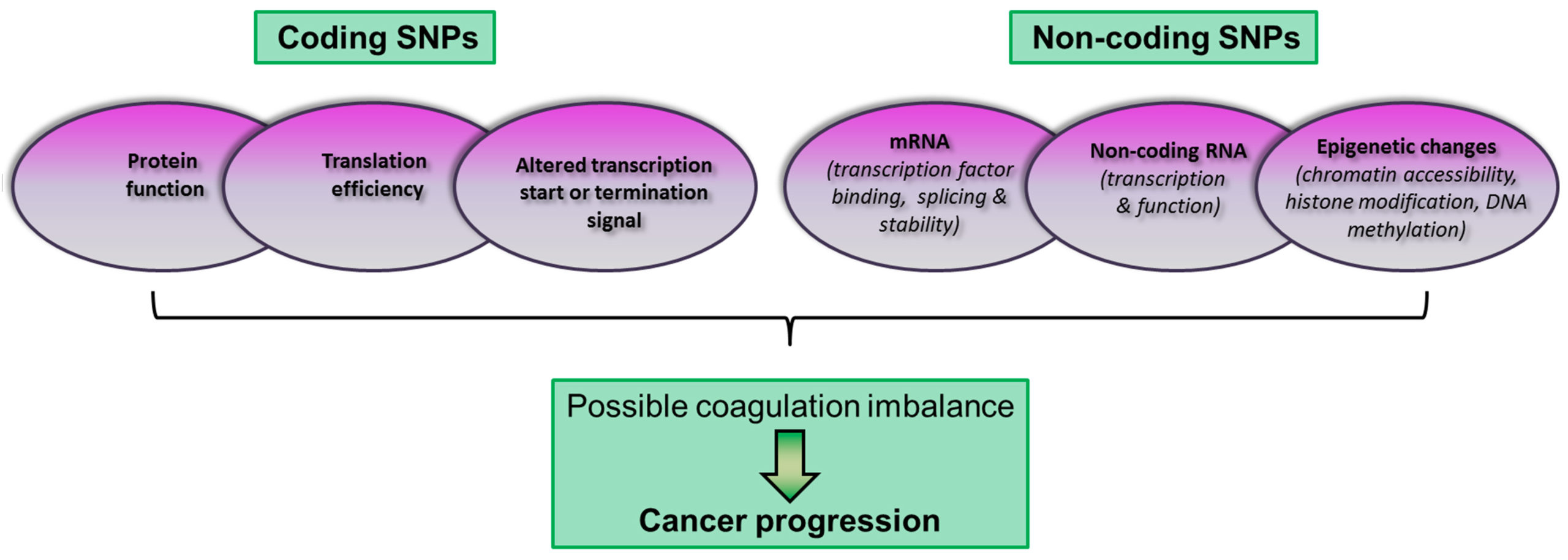
SNPs are found in both coding and non-coding regions and they may have functional effects relevant to disease phenotypes (Fig. 2).
It is now believed that true disease associated SNPs exert their effects by altering gene expression. High-throughput sequencing of human genomes, transcriptomes and epigenomes have opened new horizons to understand the genetic basis of oncogenesis and revealing translational research opportunities. Hopefully, this will motivate more groups to perform research aiming to better describe the intricate relationship between cancer and coagulation at the genetic level. One initial approach could be to epigenetic markers and open chromatin regions to identify SNPs in or near coagulation genes that reside in regulatory regions, such as transcription factor binding sites. Disclosing the role of such polymorphisms in genes of the coagulation system on cancer risk and gene expression may be clinically useful and represents one means to achieve a mechanistic understanding of the hemostatic abnormalities in cancer patients. Increased knowledge within the field of cancer and coagulation has the potential to be translated into more individualized approaches for the management and prevention of cancer and cancer-associated thrombotic complications.
References
- Jiang, X., Y.L. Guo, and M.E. Bromberg, Formation of tissue factor-factor VIIa-factor Xa complex prevents apoptosis in human breast cancer cells. Thromb Haemost, 2006. 96(2): p. 196-201.
- Versteeg, H.H., F. Schaffner, M. Kerver, H.H. Petersen, J. Ahamed, B. Felding-Habermann, Y. Takada, B.M. Mueller, and W. Ruf, Inhibition of tissue factor signaling suppresses tumor growth. Blood, 2008. 111(1): p. 190-9.
- Falanga, A., L. Russo, and V. Milesi, The coagulopathy of cancer. Curr Opin Hematol, 2014. 21(5): p. 423-9.
- Stavik, B., G. Skretting, M. Sletten, P.M. Sandset, and N. Iversen, Overexpression of Both TFPI alpha and TFPI beta Induces Apoptosis and Expression of Genes Involved in the Death Receptor Pathway in Breast Cancer Cells. Mol Carcinog, 2010. 49(11): p. 951-963.
- Stavik, B., G. Skretting, H.C. Aasheim, M. Tinholt, L. Zernichow, M. Sletten, P.M. Sandset, and N. Iversen, Downregulation of TFPI in breast cancer cells induces tyrosine phosphorylation signaling and increases metastatic growth by stimulating cell motility. Bmc Cancer, 2011. 11: p. 357.
- Tinholt, M., H.K. Vollan, K.K. Sahlberg, S. Jernstrom, F. Kaveh, O.C. Lingjaerde, R. Karesen, T. Sauer, V. Kristensen, A.L. Borresen-Dale, P.M. Sandset, and N. Iversen, Tumor expression, plasma levels and genetic polymorphisms of the coagulation inhibitor TFPI are associated with clinicopathological parameters and survival in breast cancer, in contrast to the coagulation initiator TF. Breast Cancer Res, 2015. 17(1): p. 44.
- Vossen, C.Y., M. Hoffmeister, J.C. Chang-Claude, F.R. Rosendaal, and H. Brenner, Clotting factor gene polymorphisms and colorectal cancer risk. J Clin Oncol, 2011. 29(13): p. 1722-7.
- Pihusch, R., G. Danzl, M. Scholz, D. Harich, M. Pihusch, P. Lohse, and E. Hiller, Impact of thrombophilic gene mutations on thrombosis risk in patients with gastrointestinal carcinoma. Cancer, 2002. 94(12): p. 3120-6.
- Battistelli, S., M. Stefanoni, A. Genovese, A. Vittoria, R. Cappelli, and F. Roviello, Prevalence of factor V Leiden and prothrombin G20210A in patients with gastric cancer. World J Gastroenterol, 2006. 12(26): p. 4179-80.
- Paspatis, G.A., A. Sfyridaki, N. Papanikolaou, K. Triantafyllou, A. Livadiotaki, A. Kapsoritakis, and N. Lydataki, Resistance to activated protein C, factor V leiden and the prothrombin G20210A variant in patients with colorectal cancer. Pathophysiol Haemost Thromb, 2002. 32(1): p. 2-7.
- Tormene, D., P. Beltramello, M. Perlati, B. Brandolin, S. Barbar, G. De Toffoli, and P. Simioni, The risk of cancer progression in women with gynecological malignancies and thrombophilic polymorphisms: a pilot case-control study. Clin Appl Thromb Hemost, 2009. 15(5): p. 535-9.
- Vairaktaris, E., C. Yapijakis, J. Wiltfang, J. Ries, A. Vylliotis, S. Derka, S. Vasiliou, and F.W. Neukam, Are factor V and prothrombin mutations associated with increased risk of oral cancer? Anticancer Res, 2005. 25(3c): p. 2561-5.
- Vylliotis, A., C. Yapijakis, E. Nkenke, T. Nisyrios, D. Avgoustidis, M. Adamopoulou, V. Ragos, S. Vassiliou, N. Koronellos, and E. Vairaktaris, Effect of thrombosis-related gene polymorphisms upon oral cancer: a regression analysis. Anticancer Res, 2013. 33(9): p. 4033-9.
- Sciacca, F.L., E. Ciusani, A. Silvani, E. Corsini, S. Frigerio, S. Pogliani, E. Parati, D. Croci, A. Boiardi, and A. Salmaggi, Genetic and plasma markers of venous thromboembolism in patients with high grade glioma. Clin Cancer Res, 2004. 10(4): p. 1312-7.
- Eroglu, A., A. Ozturk, and N. Akar, Association between the -402GA, -401GT, and -323ins10-bp polymorphisms of factor VII gene and breast cancer. Breast Cancer, 2011. 18(4): p. 282-285.
- Tinholt, M., M.K. Viken, A.E. Dahm, H.K. Vollan, K.K. Sahlberg, O. Garred, A.L. Borresen-Dale, A.F. Jacobsen, V. Kristensen, I. Bukholm, R. Karesen, E. Schlichting, G. Skretting, B.A. Lie, P.M. Sandset, and N. Iversen, Increased coagulation activity and genetic polymorphisms in the F5, F10 and EPCR genes are associated with breast cancer: a case-control study. BMC Cancer, 2014. 14(1): p. 845.
